A short demo on how to use IPython Notebook as a research notebook
As promised, here's the IPython Notebook tutorial I mentioned in my introduction to IPython Notebook.
Downloading and installing IPython Notebook
You can download IPython Notebook with the majority of the other packages you'll need in the Anaconda Python distribution. From there, it's just a matter of running the installer, clicking Next and Accept buttons a bunch of times, and voila! IPython Notebook is installed.
Running IPython Notebook
For Mac and Linux users, open up your terminal. Windows users need to open up their Command Prompt. Change directories in the terminal (using the cd command) to the working directory where you want to store your IPython Notebook data.
To run IPython Notebook, enter the following command:
ipython notebook
It may take a minute or two to set itself up, but eventually IPython Notebook will open in your default web browser and should look something like this:
(NOTE: currently, IPython Notebook only supports Firefox and Chrome.)
Creating a new notebook
Conveniently, Titus Brown has already posted a quick demo on YouTube. (Start at 2m16s.)
Now that we've covered the basics, let's get into how to actually use all this as a research notebook.
Using IPython Notebook as a research notebook
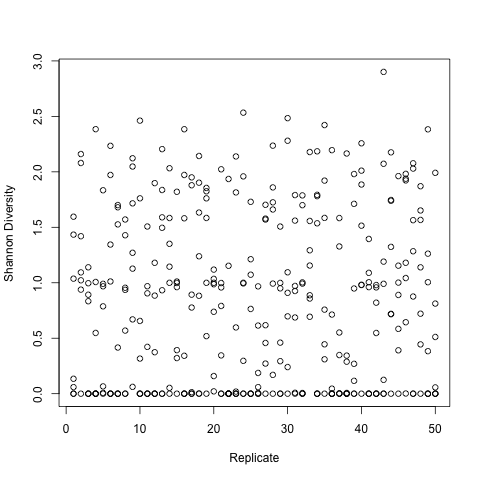
The great part about the seamless integration of text and code in IPython Notebook is that it's entirely conducive to the "form hypothesis - test hypothesis - evaluate data - form conclusion from data - repeat" process that we all follow (purposely or not) in science. For this example, let's say we're studying an Artificial Life swarm system and the effects of various environmental parameters on the swarm.
Here's the example research notebook: [pdf] [ipynb w/ accompanying files]
I designed this demo research notebook to be a self-guided tour through the thought process of a researcher as he works on a research project, so hopefully it's helpful to other researchers out there.
Statistics in IPython Notebook
UPDATE (10/19/2012): Please refer to my other blog post for an up-to-date guide on statistics in Python.
For those of you who (understandably) don't want to search through an entire research notebook to figure out how to do statistics in IPython Notebook, here's the cut and dry code.
Reading data
control1 is the dictionary of parsed data
control1_columns is the list of column names used to access the data dictionary, sorted in the same order as the csv data file.
NOTE: This uses a function from my custom Python library, which parses the data into convenient data dictionaries.
The data in the dictionaries can be accessed by:
Standard error of the mean
Bootstrapped 95% confidence intervals
The code below shows you how to compute bootstrapped 95% CIs for the mean. However, this function can bootstrap any range of CIs for any statistical function (mean, mode, standard deviation, etc.). Here's the input parameter description:
Input parameters: data = data to get bootstrapped CIs for statfun = function to compute CIs over (usually, mean) alpha = size of CIs (0.05 --> 95% CIs). default = 0.05 n_samples = # of bootstrap populations to construct. default = 10,000 Returns: bootstrapped confidence intervals, formatted for the matplotlib errorbar() function
NOTE: This uses a couple functions from my custom Python library, since bootstrapping CIs isn't currently supported by SciPy/NumPy.
Mann-Whitney-Wilcoxon RankSum test
Analysis of variance (ANOVA)
SciPy's ANOVA function takes two or more dataset lists as its input parameters.
Hopefully everyone finds this useful. Get in touch if you have any more ideas on IPython Notebook as a research notebook, or if you'd like to figure out how to do some more statistical tests in Python.

Dr. Randal S. Olson
AI Researcher & Builder · Co-Founder & CTO at Goodeye Labs
I turn ambitious AI ideas into business wins, bridging the gap between technical promise and real-world impact.